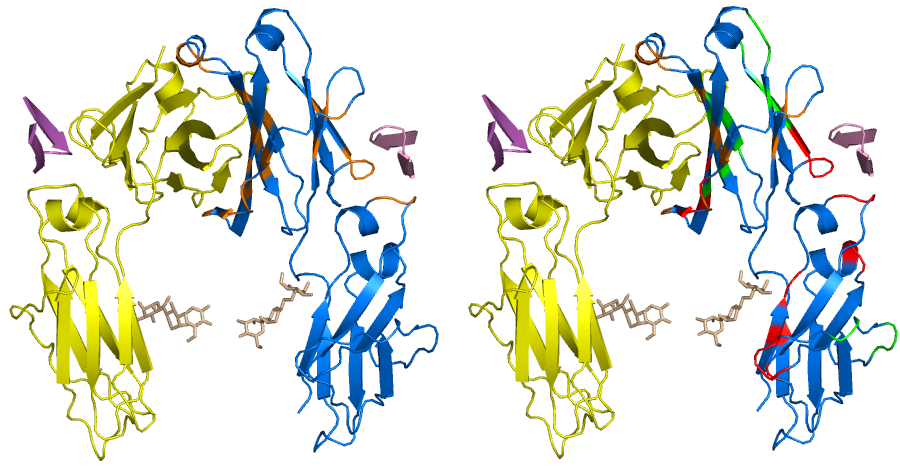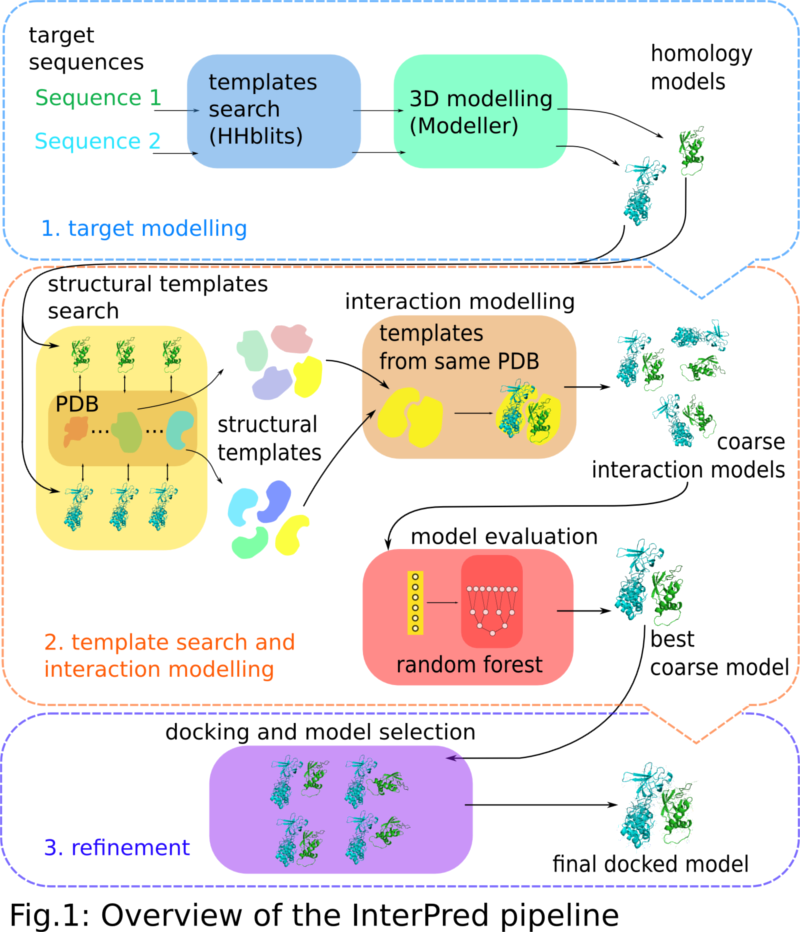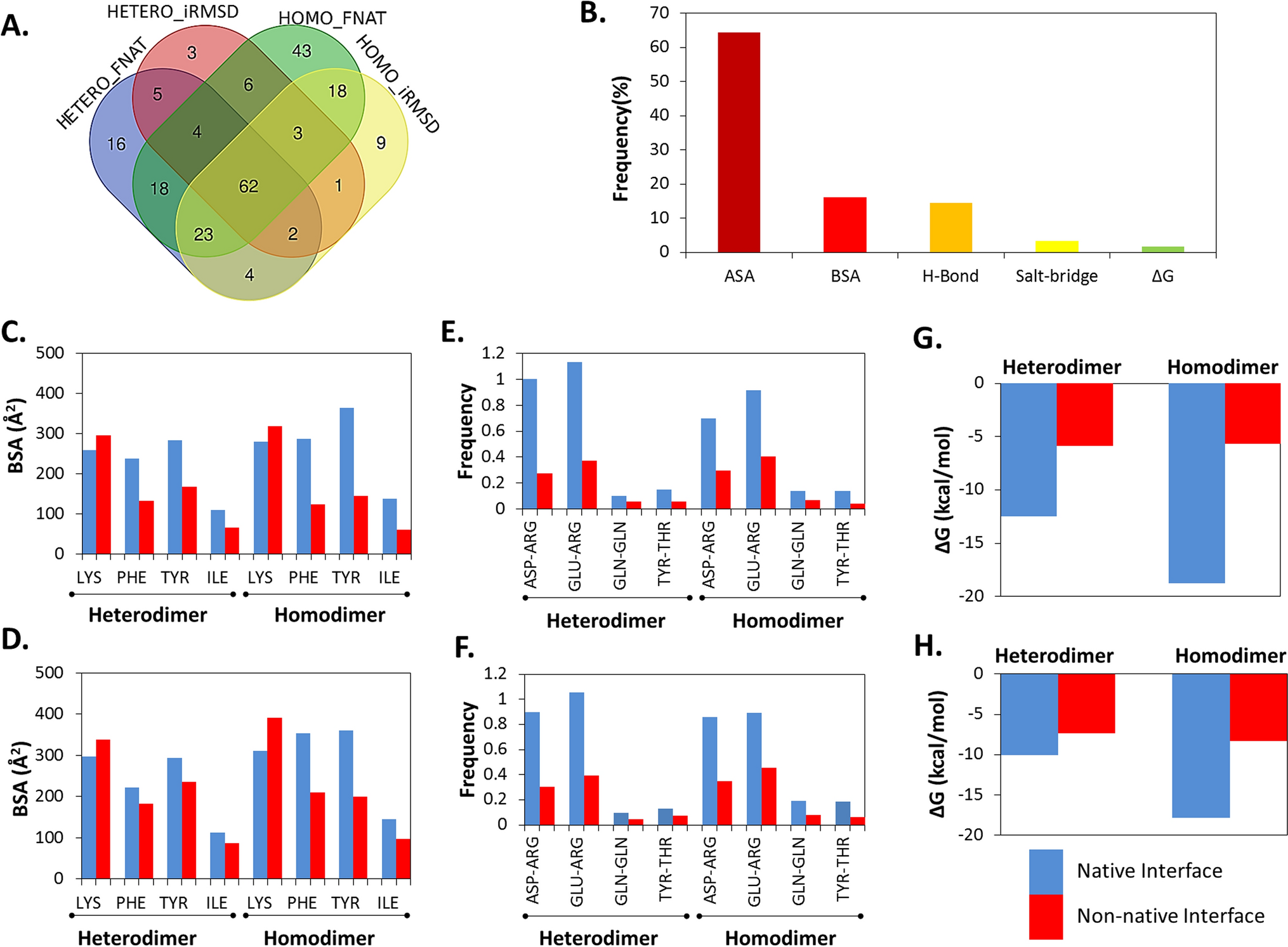
Classification and prediction of protein–protein interaction interface using machine learning algorithm | Scientific Reports
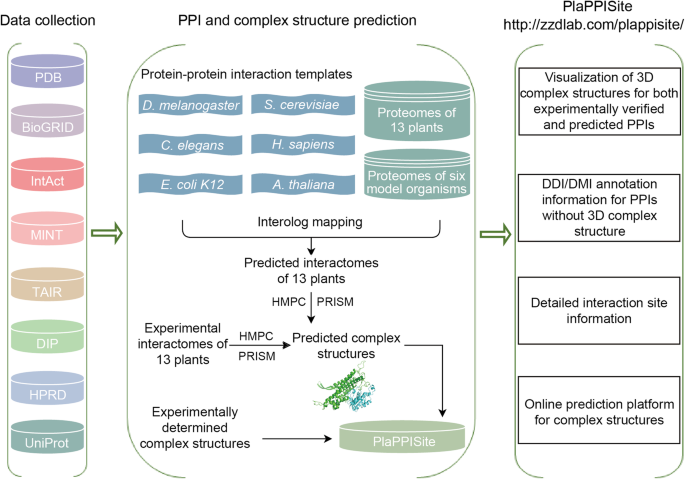
PlaPPISite: a comprehensive resource for plant protein-protein interaction sites | BMC Plant Biology | Full Text
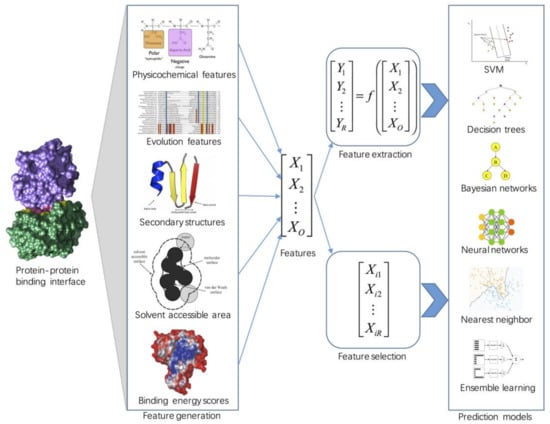
Molecules | Free Full-Text | Machine Learning Approaches for Protein–Protein Interaction Hot Spot Prediction: Progress and Comparative Assessment | HTML

Protein ^ protein interaction prediction. PPIs can be predicted based... | Download Scientific Diagram
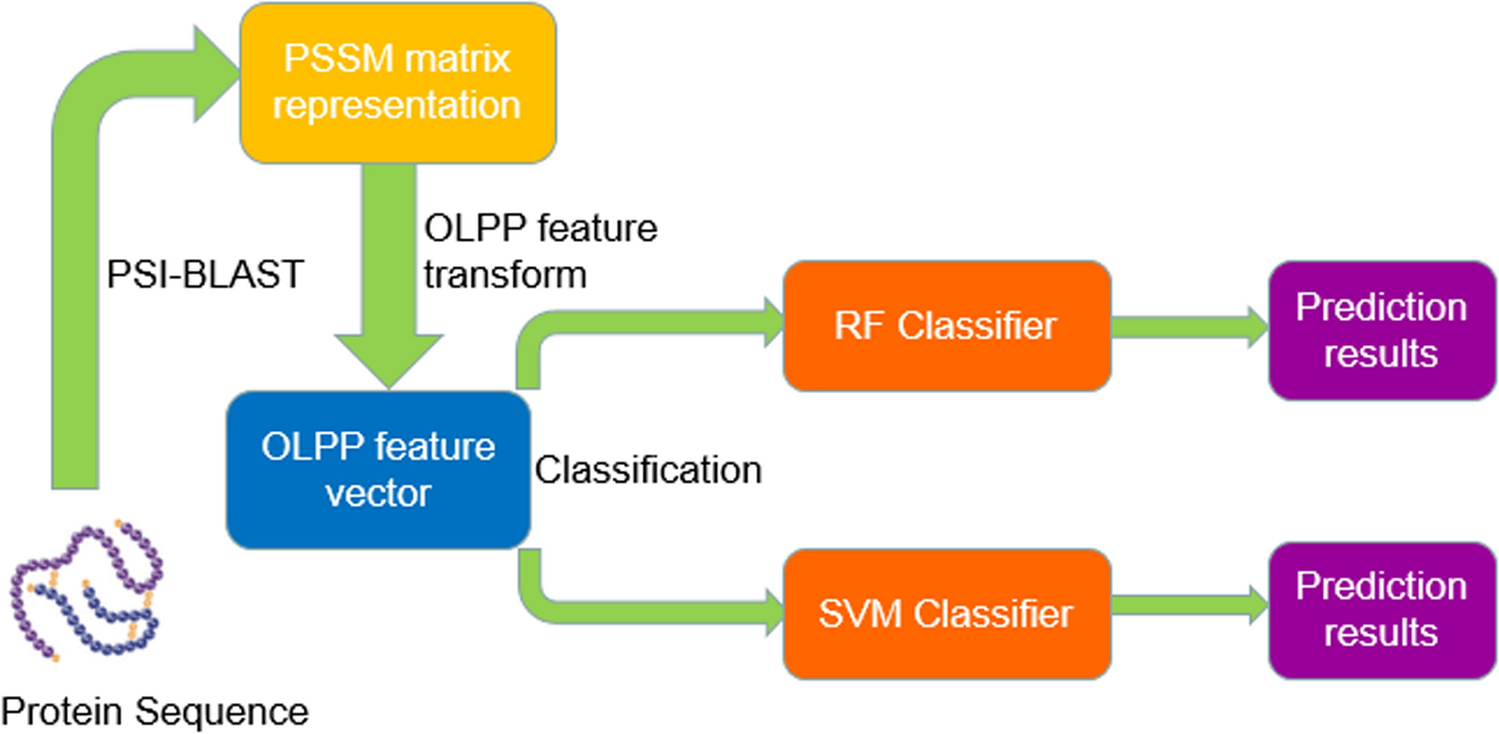
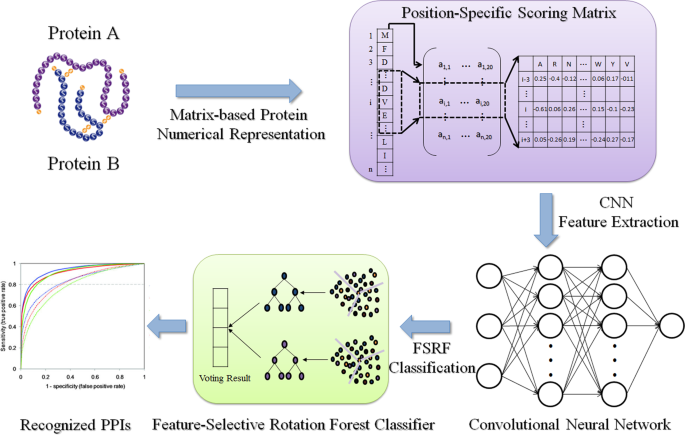
Robust and accurate prediction of protein–protein interactions by exploiting evolutionary information | Scientific Reports
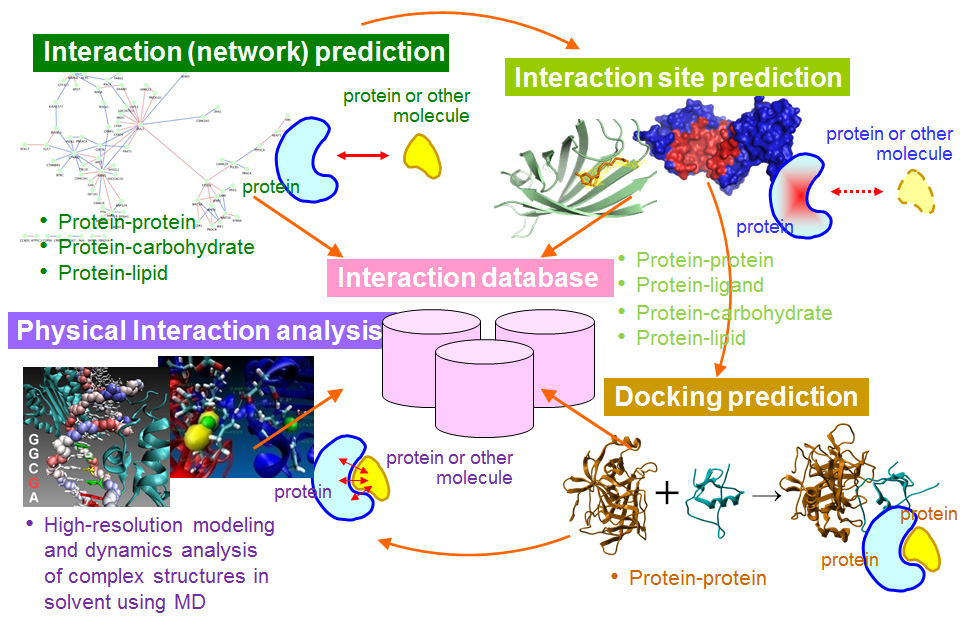
Plantae | Proteome-wide, structure-based prediction of protein-protein interactions (Plant Physiol) | Plantae
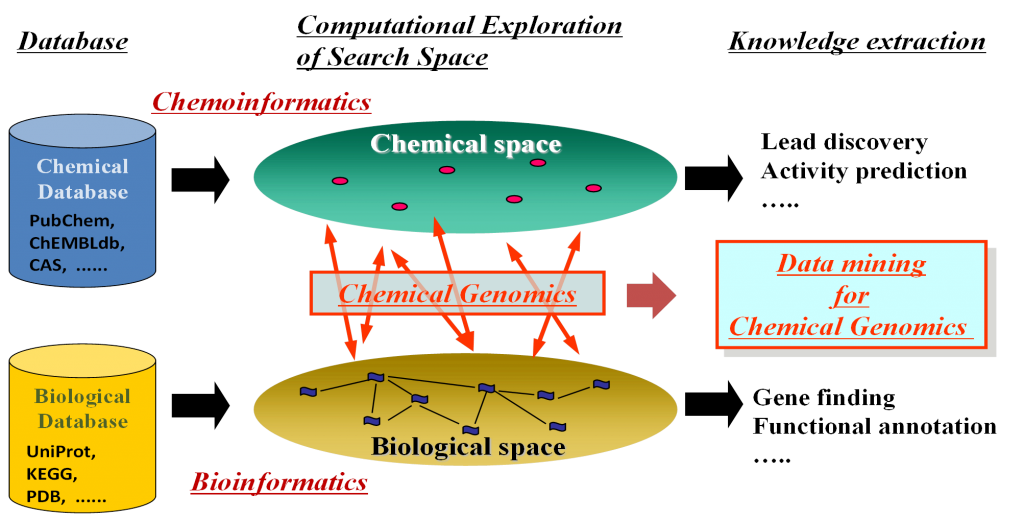
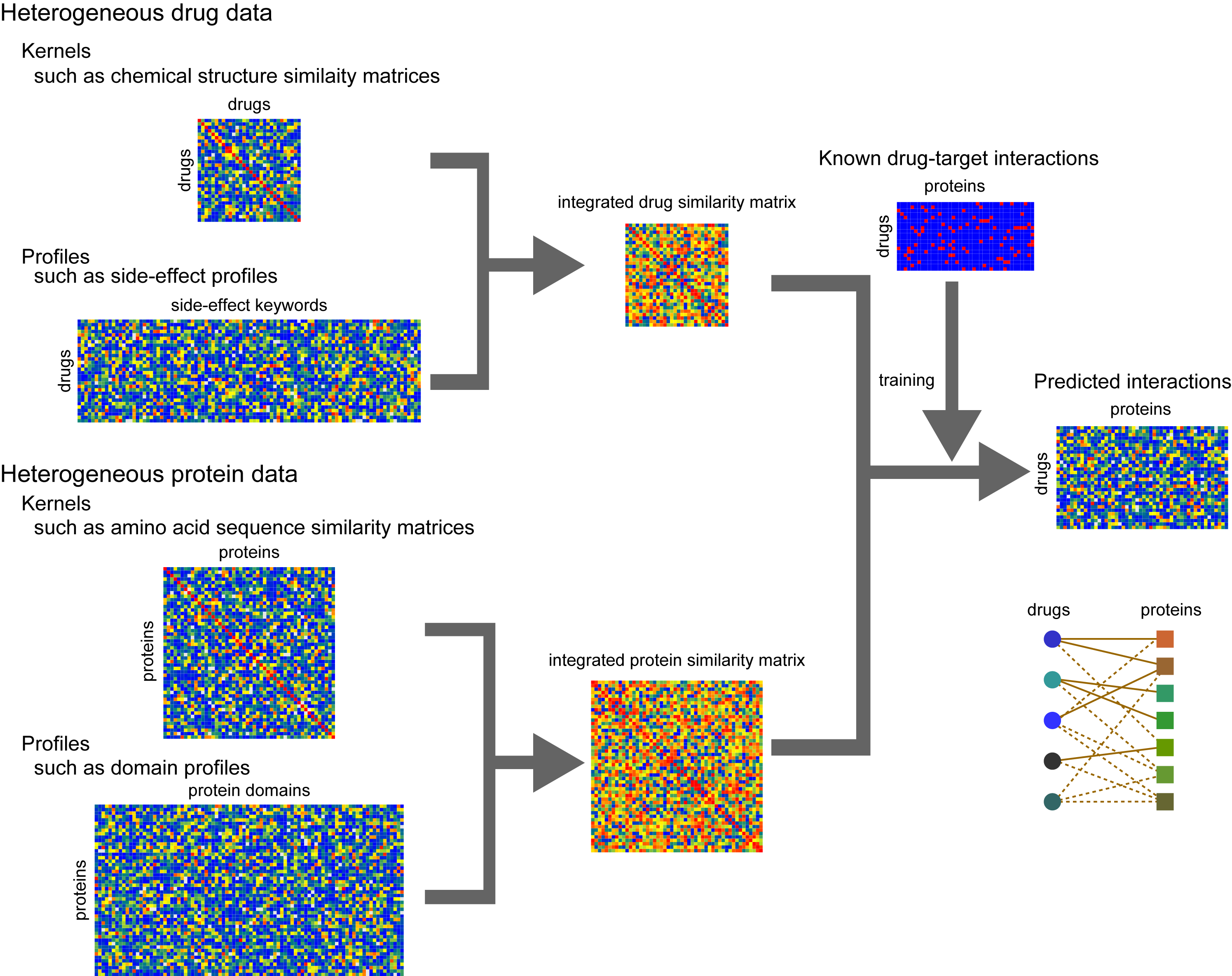
In silico target prediction for protein-protein interaction (PPI) small molecule inhibitors using CGBVS - In Silico 創薬

Protein-assisted RNA fragment docking (RnaX) for modeling RNA–protein interactions using ModelX | PNAS

IJMS | Free Full-Text | Prediction of Protein–Protein Interactions by Evidence Combining Methods | HTML
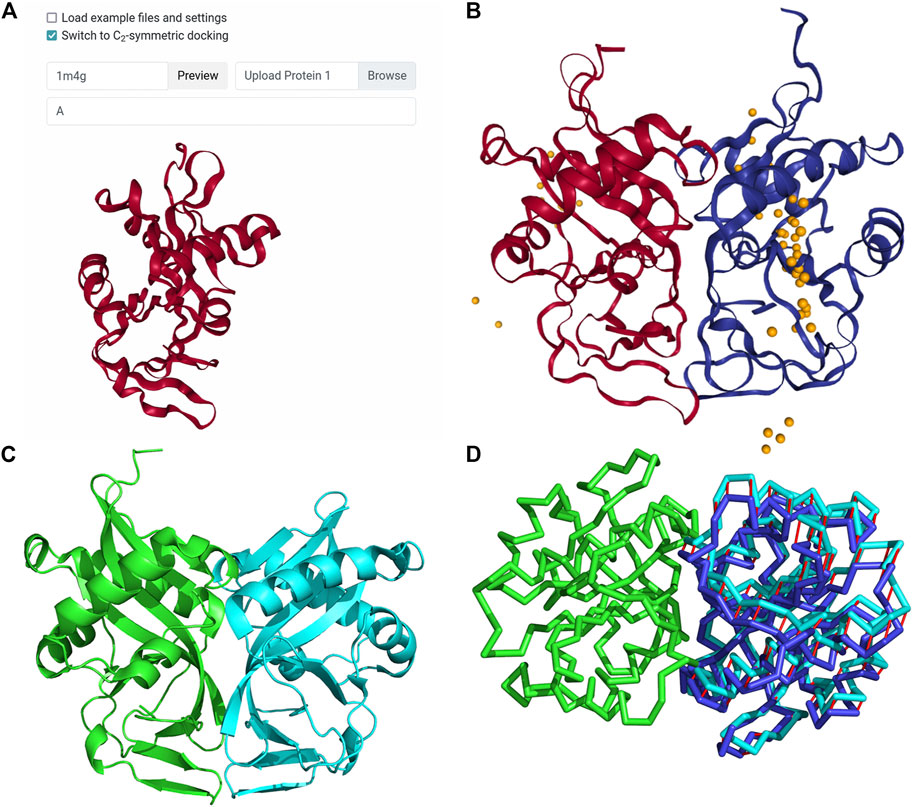
Frontiers | LZerD Protein-Protein Docking Webserver Enhanced With de novo Structure Prediction | Molecular Biosciences
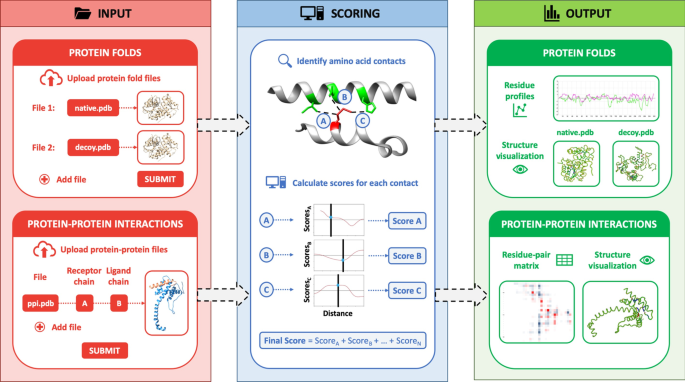
SPServer: split-statistical potentials for the analysis of protein structures and protein–protein interactions | BMC Bioinformatics | Full Text

BindML: Software for Predicting Protein-Protein Interaction Sites using Phylogenetic Substitution Models

BindML: Software for Predicting Protein-Protein Interaction Sites using Phylogenetic Substitution Models

Scheme for all-to-all protein-protein interaction predictions. Using a... | Download Scientific Diagram
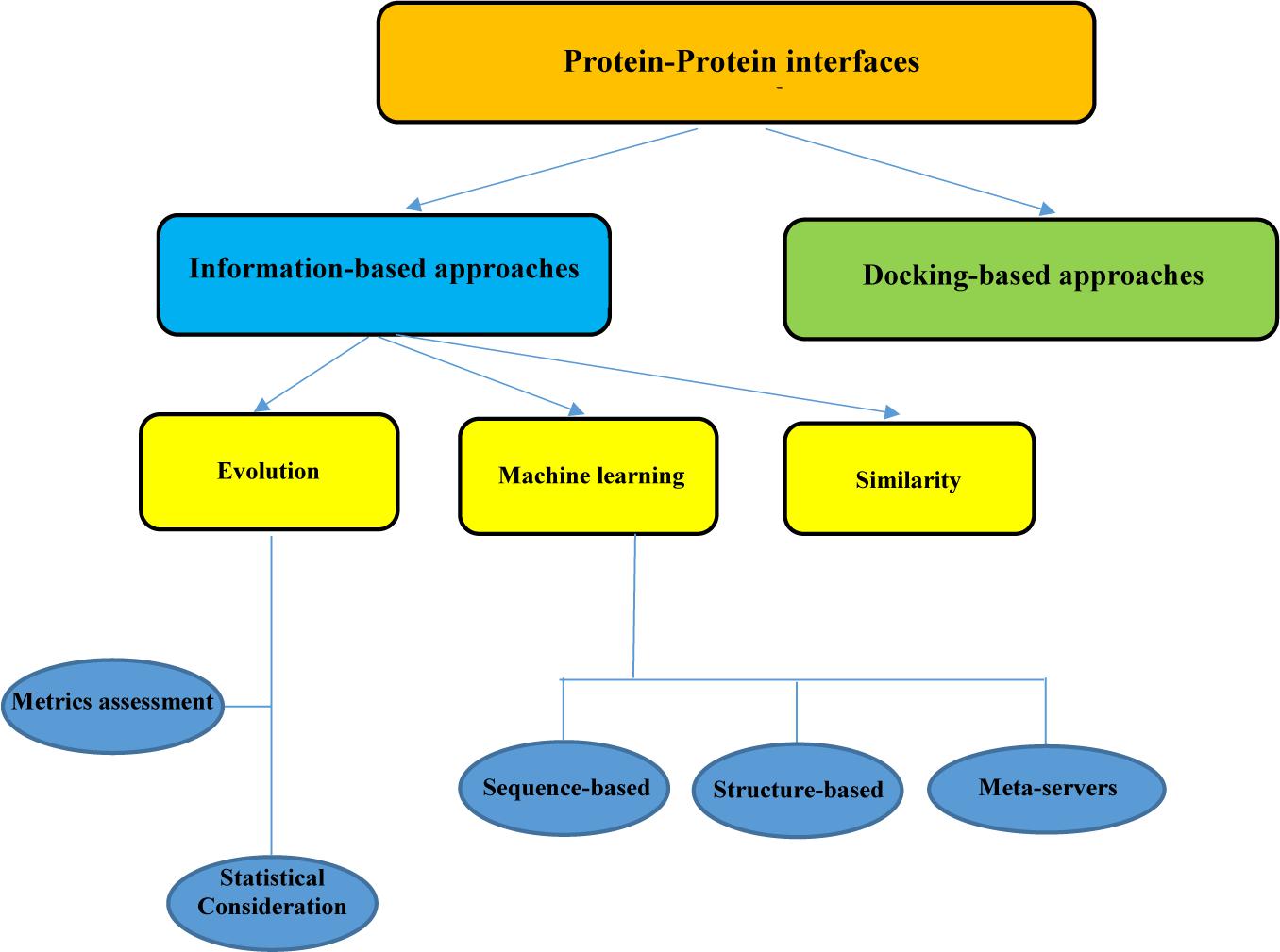
Frontiers | In silico Approaches for the Design and Optimization of Interfering Peptides Against Protein–Protein Interactions | Molecular Biosciences